Example: training and test sets
- sTrainD.data: numerical measurement values for the training samples, 4 measurements /individual
- sTrainD.label: true class labels of the training set: 'Setosa','Versicolor' or 'Virginica'
- sTestD.data: numerical measurement values for the test samples, 4 measurements /individual
- sTestD.label: true class labels of the test set: 'Setosa','Versicolor' or 'Virginica'
Let's examine only one of the measurements as an example.
Below is a plot of the values of training set variable
'Sepal width' (sTrainD.data(:,2)).
For the training set, the true class corresponding to each
value is known. It can be observed that
the species 'Setosa' has somewhat wider sepal than the
two othe species.
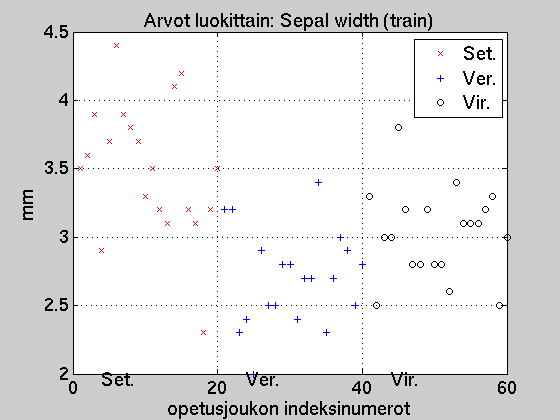
On the basis of this analysis, we can state that if the sepal width
is larger than 3.4 mm the sample is
a 'Setosa' with a rather large probability.
If we received new measurements of sepal widths,
we could draw them as below.
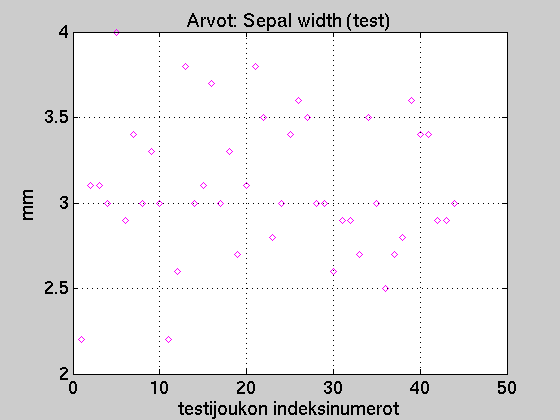
Our task would be to guess/decide the species of the new samples on basis of
the sepal withs only.
On the basis of the above analysis, we could classify all
the samples with sepal with over 3.4 mm as 'Setosa'.
Probably we would make classification errors:
we would classify 'Virginica' and 'Versicolor' samples as 'Setosas'.
In order to assess the quality of the classifier we use,
the data set is usually partitioned into (at least) two, training and test
sets.
By knowing the correct labels also for the test set it is straightforward
to compute how many of the proposed classifications were correct.
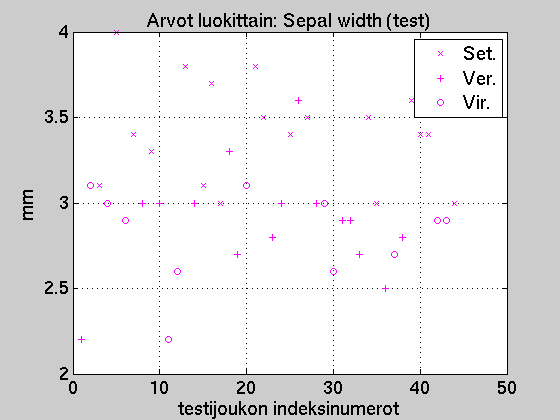
If all the samples with sepal width over 3.4mm
were classified as
'Setosa', at least one 'Versicolor' (index 26)
would be misclassified
NOTE! In the data set, there are four
different measurements and all of them are used jointly.
The classification is performed primarily by the KNN
algorithm.


